EIS Analyzer - Professional Impedance Analysis
Electrochemical impedance spectroscopy (EIS) analysis software for battery impedance testing, fuel cell diagnostics, and electrochemical R&D. Features automated circuit fitting, DRT analysis, Kramers-Kronig validation, and Batalyse Collect integration. Free for Data Analysis customers.

EIS Analyzer

Multiple File Formats & Data Sources
Import EIS data from Gamry (.DTA), Solartron (.z), Bio-Logic (.mpr), PalmSens (.pssession), or text formats (.txt, .dat, .csv, .mpt). Connect directly to MySQL/SQL Server databases or Batalyse Collect cloud platform.
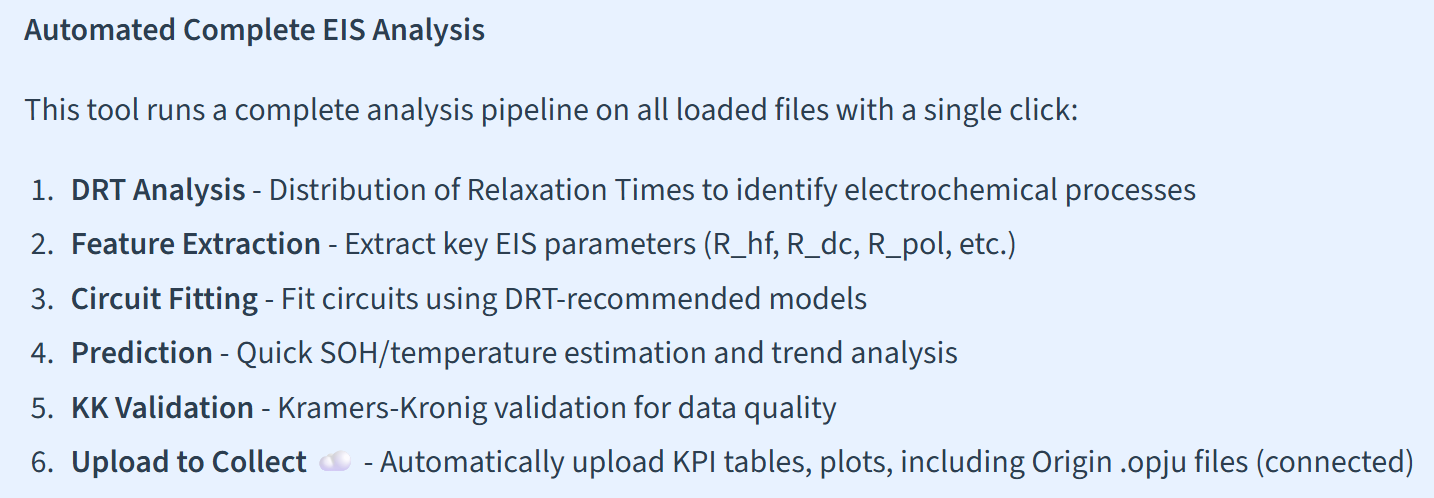

One-Click Automated Analysis
Run DRT analysis, circuit fitting, validation, and predictions simultaneously with the auto-analysis pipeline. Export results as JSON, CSV, or PNG – or upload directly to Collect with KPI tables.

Customization
Customize every aspect of your graph exports with the dedicated Customize Graphs tab. Set per-file dataset styling including colors, markers, and line widths. Configure font settings with bold and italic options for titles, axis labels, and legends. Control axis scaling, gridlines, and select which plots to generate. Save and share your styling presets via JSON export/import.
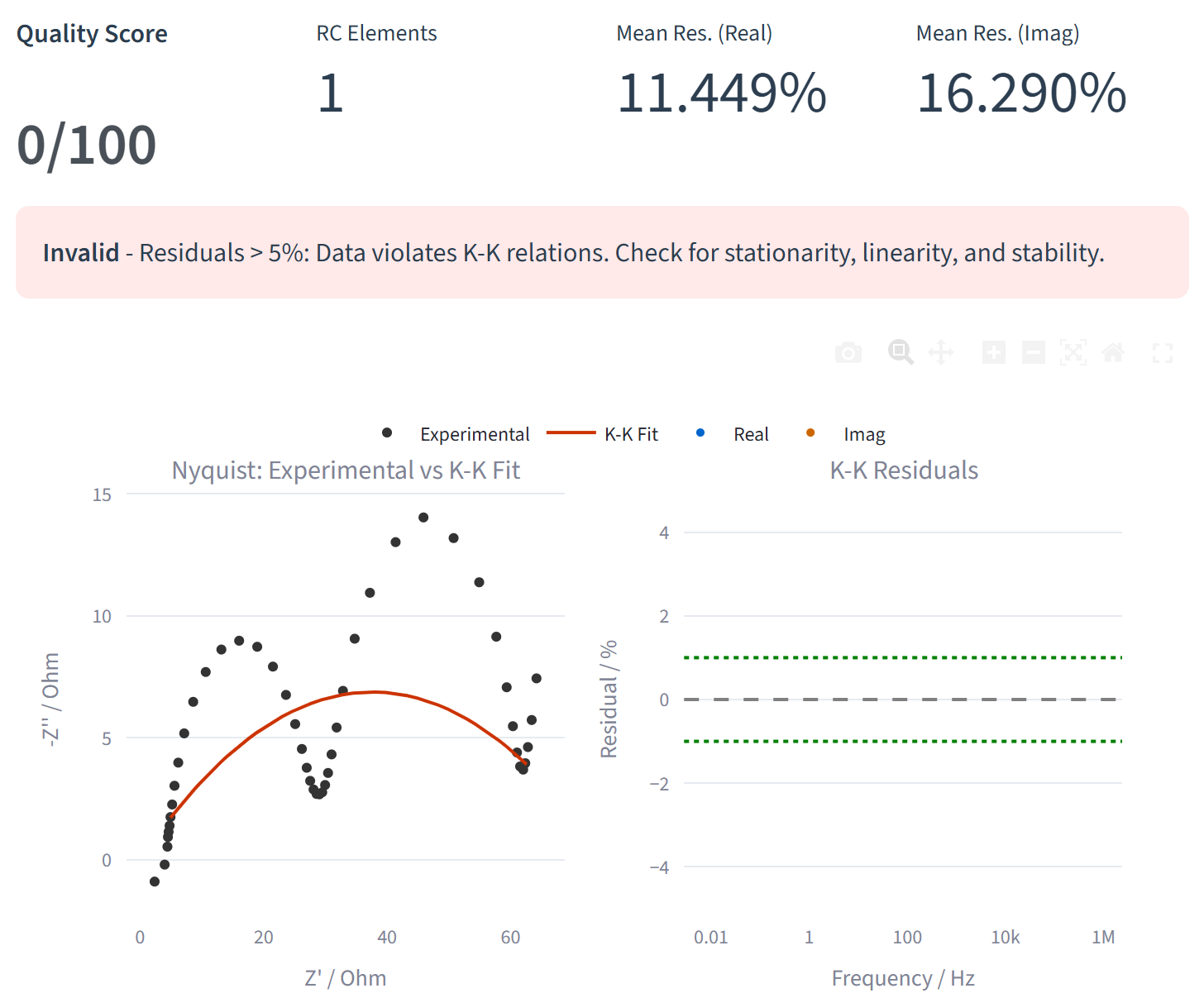
Kramers-Kronig Validation
Verify the quality of your EIS measurements with Kramers-Kronig transformation. Detect drift, noise, or non-linear behavior in your data before analysis. The validation highlights problematic frequency ranges and helps ensure reliable circuit fitting results.

Distribution of Relaxation Times
Identify electrochemical processes and their time constants with Distribution of Relaxation Times analysis. Extended configuration options let you fine-tune regularization parameters, frequency ranges, and peak detection sensitivity. Add peaks manually by clicking directly on the DRT graph or let the algorithm auto-detect them. DRT results automatically suggest optimal equivalent circuit models for subsequent fitting.

Equivalent Circuit Modeling
Fit your EIS data to 14+ built-in equivalent circuit models including R, R-C, R-RC, R-RQ, R-RC-RC, R-RQ-RQ, Randles, and models with Warburg diffusion elements. Need a custom configuration? Enter your own circuit string to define individual equivalent circuits tailored to your specific electrochemical system.

Save and manage projects
Save complete analysis sessions as projects to preserve all settings, fitted parameters, and results. Reopen projects anytime to continue your work or compare results across different measurements.

Flexible Export Options
Export your analysis results in multiple formats: JSON for programmatic access, CSV for spreadsheet analysis, PNG for publication-ready graphs, or editable OriginLab .opju projects. Upload results directly to Batalyse Collect for centralized data management and team collaboration.
Free for Data Analysis Customers
Changelog
- New Customization Library for saving and loading graph presets
- Position-based defaults for single-test presets in Customization Library
- DRT Analysis: Added multiple new configuration options
- DRT Analysis: Add peaks manually or by clicking on the graph
- Auto Analysis: DRT settings can now be modified before running
- Added DRT peak information to Origin export with Peak_type column
- Auto-extract DRT peaks when running Auto Analysis and exporting
- Added Test X: prefix to Styles menu for clarity
- Fixed bugs when loading parameters from presets
- Improved default column tooltip in Customization Library
- Bug fixes and performance improvements
- Fixed startup crash when installed in protected directories (Program Files)
- Application log now writes to user directory instead of install directory
- Added SQL injection protection for custom database query configurations
- Created comprehensive security documentation for IT departments
- Fixed Solartron ZPLOT2 .z file parsing (extended column format support)
- Added Customize Graphs tab with full graph export customization
- Added per-file dataset styling (color, marker, line width)
- Added bold/italic font settings for titles, axis labels, legends
- Added JSON customization export/import for sharing settings
- Added export graphs selection to control which plots are generated
- Added Nyquist_Areal graph for area-normalized impedance plots
- Added PalmSens .pssession file format support
- Fixed combined Origin export for multi-file graphs
- Improved fitting failure messages with model name and error details
Version 1.0.5
- Added Origin export with KK_Nyquist graph (Experimental vs K-K Fit)
- Improved Origin detection with registry scanning and refresh button
- Fixed Kramers-Kronig validation to use proper linKK function
- Enhanced KK validation error display with residual data and plots
- Fixed local graph export (PNG ZIP) to include Nyquist and Bode plots
- Added terms acceptance documentation to license requests
- Fixed Origin uploads to Collect
Version 1.0.4
- Added Fitting Options expander to Auto Analysis with configurable parameters
- Improved MPR parser to handle high-impedance data (raised filter from 1kΩ to 1MΩ)
- Fixed UI reloads when modifying fitting options using Streamlit fragments
Version 1.0.3
- Added combined DRT comparison plot for multi-file exports
- Added combined KK validation plot for multi-file exports
- Combined Nyquist/Bode plots now generated automatically for comparison uploads
- Online licenses now re-validate on each app startup
- Enhanced offline license activation with key decryption capability
Version 1.0.2
- Fixed license validation on app startup and browser refresh
- Resolved tab-switching issue when interacting with checkboxes
- Bug Report tab now accessible without valid license
Version 1.0.1
- Replaced pyodbc with mssql-python for SQL Server connectivity
- Added sidebar expand button when collapsed
- Made Origin packages Windows-only to fix install errors
Credits

- EIS-Data-Analytics
https://github.com/isea-rwth-aachen/EIS-Data-Analytics
Blömeke, A., Kappelhoff, O., & Sauer, D. U. (2024). EIS Data Analytics. - PyEIS
https://github.com/kbknudsen/PyEIS
Knudsen, K. B. - impedance.py
https://github.com/ECSHackWeek/impedance.py
Murbach, M. D., Gerwe, B., Dawson-Elli, N., & Tsui, L. - pyDRTtools
https://github.com/ciuccislab/pyDRTtools
Ciucci, F., Chen, C., Effat, M. B., Liu, J., Maradesa, A., Py, B., Saccoccio, M., & Wan, T. H.
